AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
 
At first, protein contact map is used to predict whether the Euclidean distance between two C β atoms is less than 8.0 Å ( Hanson et al., 2018 Wang et al., 2017). Protein contact map is critical to template-free modeling. Our previous work OPUS-TASS ( Xu et al., 2020c) introduced some new features derived from our potential functions ( Lu et al., 2008 Xu et al., 2017, 2018) to improve the accuracy. Many successful methods have been proposed to predict protein 1D features ( Fang et al., 2018 Gao et al., 2018 Hanson et al., 2019 Heffernan et al., 2017 Klausen et al., 2019 Xu et al., 2020c), among which SPIDER3 ( Heffernan et al., 2017) and NetSurfP-2.0 ( Klausen et al., 2019) adopted bidirectional recurrent neural networks to measure long-range interactions, SPOT-1D ( Hanson et al., 2019) integrated the predicted contact map ( Hanson et al., 2018) to capture protein global information. 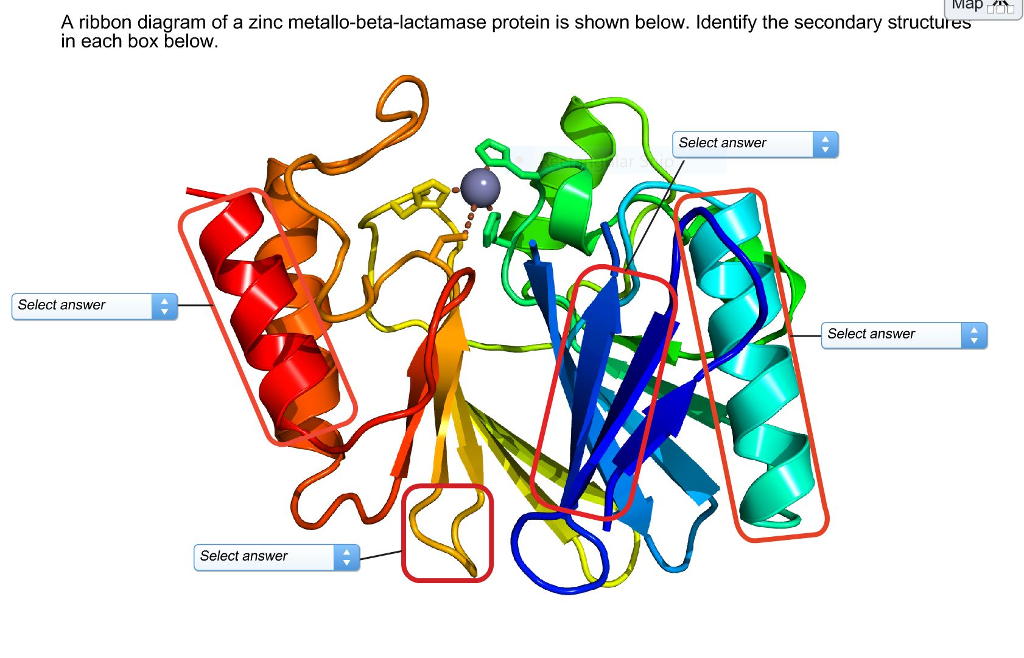
Protein solvent accessibility measures the residue’s exposure to solvent at its folded state. 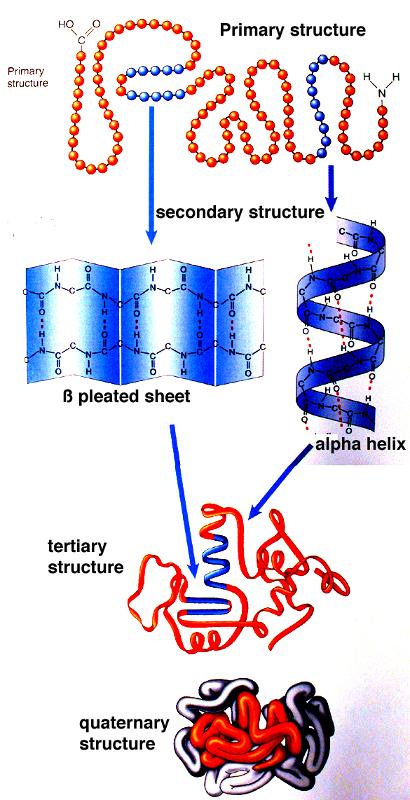
Protein secondary structure has been classified into either 3- or 8-state ( Kabsch and Sander, 1983), and it can be used to describe protein local conformation. Therefore, most researches only take Φ and Ψ into consideration ( Hanson et al., 2019 Heffernan et al., 2017 Klausen et al., 2019 Xu et al., 2020c). Among them, Ω is around 180° in most case. Protein backbone torsion angles (Φ, Ψ and Ω) determine the entire protein conformation. In the recent 14th Community-Wide Experiment on the Critical Assessment of Techniques for Protein Structure Prediction (CASP14), AlphaFold2 developed by DeepMind exhibits astonishingly performance ( Jumper et al., 2020), indicating that the computational methods have reached a practicable level.Īlthough protein 3D structure prediction is important, there are scenarios in which high-accuracy prediction of low-dimensional structural features, such as 1D features like torsion angles (Φ and Ψ), secondary structure (3-state and 8-state) and solvent accessibility, may be useful for successive modeling ( Xu et al., 2020a). 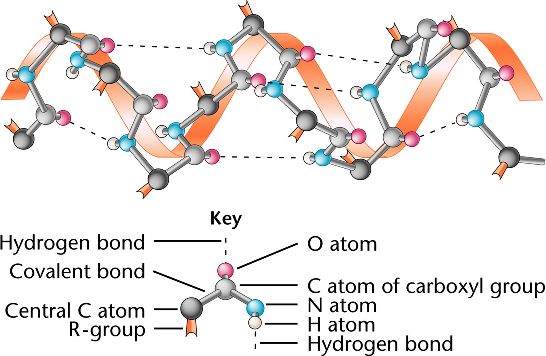
In recent years, with the development of deep learning techniques, many methods have been proposed ( Jumper et al., 2020 Senior et al., 2020 Song et al., 2013 Wang et al., 2017 Wu et al., 2021 Yang et al., 2020 Yang and Zhang, 2015), improving the performance of protein structure prediction by a large margin. Protein 3D structure prediction is crucial since the experimental approaches are usually time-consuming. 
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
